|

Phenotypic drug discovery is successful strategy that allows to identify substances (small molecules, peptides etc.) which alter the phenotype of cells, tissues or whole organisms in a desired manner.
OTAVAchemicals offers a high quality Phenotypic Screening Library (5000 compounds in total) for such empirical approaches. This library contains a set of compounds with maximal biological and chemical diversity (more than 1000 chemical scaffolds including nature-like).
Given the potential applications of a Phenotypic Screening Library, the focus of the compounds selection strategy lies on biodiversity, screening-compliant physicochemical properties and maximal coverage of chemical space, aimed at providing hits for a wide spectrum of biological goals. The library has been designed by clustering of biologically relevant chemical database (50,000 small organic compounds). The database contained compounds selected on the base of approved drugs templates and templates of ligands with known activity against main classes of biological targets taken from ChEMBL and also included compounds from OTAVAchemicals targeted libraries.
Approved drugs were taken from DrugBank and clustered to several sets served as templates for Bayesian models. Also it was used sets of bioactive reference molecules (activity cutoff was 10 nM, only for the most numerous groups – 1 nM and for small groups – 100 nM) towards major subclasses of protein target classes (enzymes, membrane receptors, ion channels, transporters, transcription factors, proteins involves in adhesion, auxiliary transport proteins, epigenetic regulators, secreted proteins, structural proteins, surface antigens, other proteins and unclassified proteins) taken from ChEMBL. The models were developed for each training sets. They were based on FCFP4, ECFP4, FCFP6 and ECFP6 fingerprints. Molecular descriptors, such as LogP, molecular weight, number of rings, number of rotatable bonds, number of hydrogen acceptors and donors and molecular polar surface area were also involved in the construction of the models for more accuracy. At the final stage, OTAVAchemicals compound сollection was screened against these models and top-scored compounds were selected.
OTAVAchemicals targeted compound libraries included to the biologically relevant chemical database were developed earlier for different target classes (e.g., kinases, GPCRs, epigenetic targets, proteases, nuclear receptors, ion channels, etc.), inhibitors of protein-protein interaction, RNA binding compounds, peptidomimetics, glycomimetics and other.
Finally biologically relevant chemical database was clustered based on fingerprints and molecular descriptors.
The Phenotypic Screening Library represents a multi-purpose diversity set of compounds for different goals such as cellular assays, bacterial assays, tissue assays, animal models, screening against various molecular targets including with unknown structures, multiple biological targets, study of cellular processes, signaling pathways, protein-protein interactions etc.
All compounds are in stock, cherry-picking is available.
The library can be expanded or narrowed based on physicochemical and other properties.
The Phenotypic Screening Library (DB, SD, XLS, PDF format) as well as the price-list is available on request. Feel free to contact us or use on-line form below to send an inquiry if you are interested to obtain this library or if you need more information.
References:
-
Anne Mai Wassermann, Luiz M. Camargo and Douglas S. Auld. Composition and applications of focus libraries to phenotypic assays. Experimental Pharmacology and Drug Discovery, Vol. 5, Article 164, July 2014, pp. 1 - 12.
-
DC Swinney. Phenotypic vs. Target-Based Drug Discovery for First-in-Class Medicines. Clinical pharmacology & Therapeutics, Vol. 93, No. 4, April 2013, pp. 299 - 301.
The summary of the Phenotypic Screening Library characteristics:
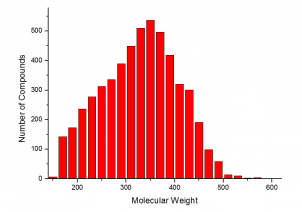  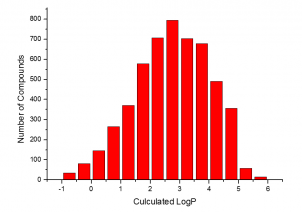  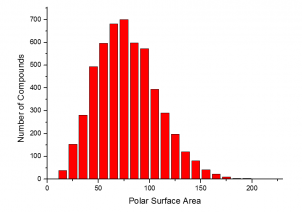 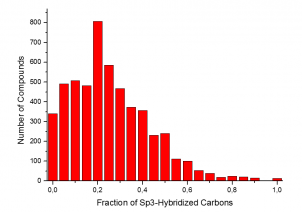 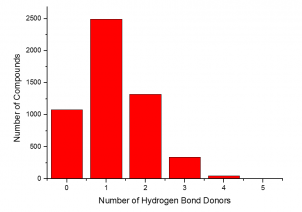 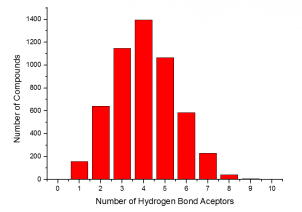  
|
 HOME
HOME ABOUT
ABOUT
 SERVICES
SERVICES
 PRODUCTS
PRODUCTS
 Targeted Libraries
Targeted Libraries
 Biochemicals
Biochemicals
 RESEARCH
RESEARCH
 DOWNLOADS
DOWNLOADS ORDERING
ORDERING
 CONTACTS
CONTACTS












